Recently, the 100th anniversary of the birth of the famous chemist Derek Barton was celebrated with a symposium. One of the many wonderful talks presented was by Tobias Ritter and entitled “Late-stage fluorination for PET imaging” and this resonated for me. The challenge is how to produce C-F bonds under mild conditions quickly so that 18F-labelled substrates can be injected for the PET imaging. Ritter has several recent articles on this theme which you should read.[cite]10.1021/ja2048072[/cite],[cite]10.1021/acs.accounts.7b00413[/cite].
The resonance was that back in 1999 I had been collaborating with colleagues to study the mechanism of rapid fluoridation using R2IF reagents.[cite]10.1039/A906212B[/cite]. This article, by the way, contains a very early example of the use of FAIR data (see here).‡ A further resonance is that Ritter computes that the displacement of an aryl-O bond by a nucleophilic fluorine is a concerted process, unlike the stepwise Meisenheimer like complexes normally occurring in nucleophilic aromatic substitutions. A few years back I explored the possibility of concerted nucleophilic substitutions, finding that F in particular was very prone to such behaviour. So it is nice to see Ritter’s real-world example of such a mechanism and indeed that his reagents (PhenoFluor) represent a significant improvement on the R2IF ones we had been exploring.
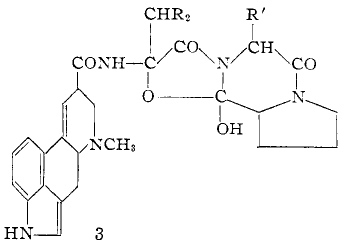
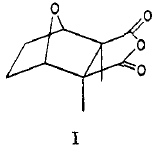
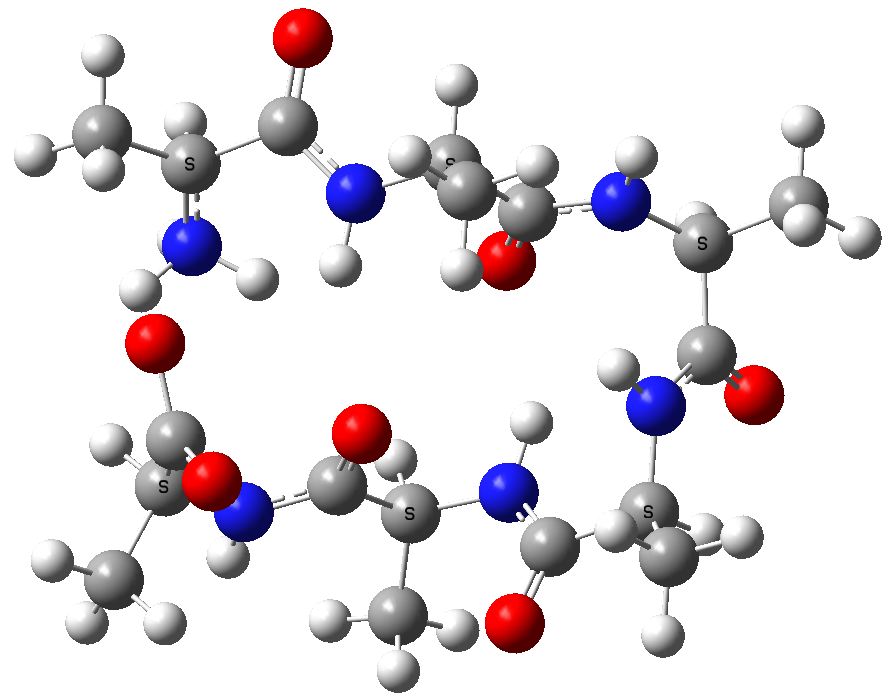
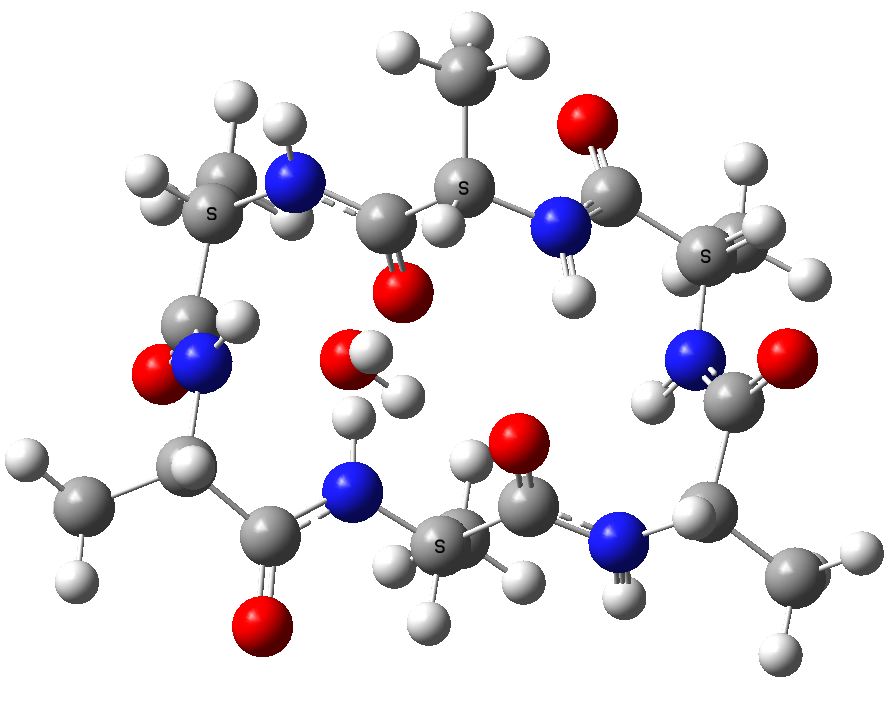
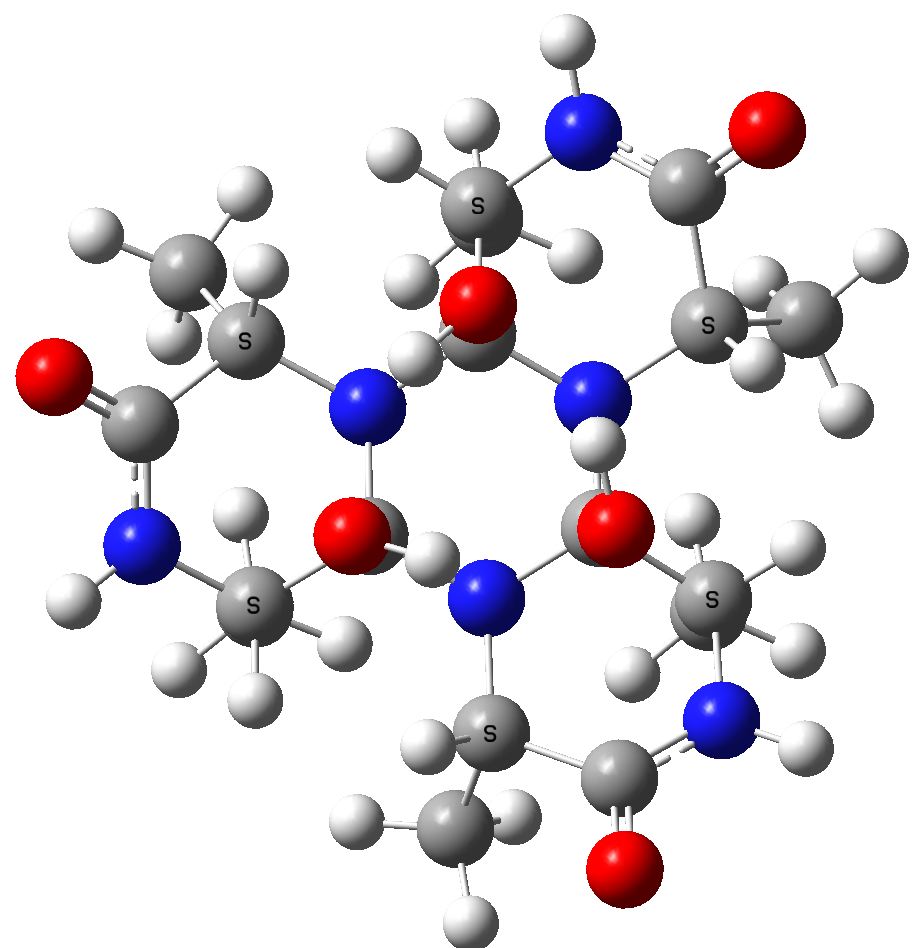
To celebrate this new chemistry, I include some results of my own which augment Ritter’s. Firstly I should start with the structure of the reagent, which contains a carbon surrounded by four heteroatoms. There are few such motifs known.♥ Thus a carbon attached to two N, one F and one O has no reported crystal structures. Relaxing the criterion to two N, one F and one other offers 71 examples, of which the most interesting are the outliers with C-F distances > 1.4Å.

The one with the value 1.5Å (DOI: 10.5517/ccdc.csd.cc1njjg5) is probably an error, as is the one at 1.45Å[cite]10.1107/S0108270189005044[/cite]. So it was a surprise to find that the calculated structure of PhenoFluor (R=2,6-di-isopropyl, B3LYP+D3BJ/Def2-TZVPPD/SCRF=toluene) had a C-F distance of 1.456Å which is surely a candidate for the longest known C(sp3)-F bond.† The computed Wiberg C-F bond order is 0.687, which is well reduced from a single bond order. This is probably due to strong anomeric effects from both nitrogens and the one oxygen, which “gang up” on the fluorine to weaken its bond and expel it as a nascent fluoride anion. Thus the E(2) NBO interaction energy is 25.1 kcal/mol for the N(Lp)-CFσ* interaction, which is unusually large, whereas the N(Lp)-COσ* interaction is only 9.2 kcal/mol.
The fluoridation is indeed computed as a concerted process, the IRC animation being shown below. Note that the trajectory of the F is initially away from the carbon but not towards the aryl group. Here it is simply forming a “hidden” fluoride anion intermediate. The trajectory then changes direction to attack the ipso-carbon. So it is a concerted but two-stage reaction path.
The IRC energy profile corresponds to a free energy barrier of 23 kcal/mol, as reported by Ritter.
Here is a less well used property along the reaction path, the dipole moment response. This shows a very abrupt charge separation in the region of the transition state and its collapse shortly after which suggests that the reaction barrier might be sensitive to the polarity of the solvent.
The issue then arises as to how much aromatic resonance is lost at the transition state. A NICS(1) aromaticity probe placed 1Å above the centroid of the aryl ring has the value -9.3 ppm, close to the value of ~ -10ppm for benzene itself. So this relatively facile reaction is in part due to significant preservation of the aryl stabilisations by aromatic resonance.
To close, the Phenofluor reagent is commercially available as a 0.1M solution in toluene, which makes one wonder if it is possible to obtain crystals. It might be of course that when the solution is concentrated, it reverts to the iminium fluoride ion pair shown above. But if crystals are possible, then it would be interesting to verify that the C-F bond in this species is indeed unusually long, perhaps even a record holder?
‡The data represent an early use of the Chime plugin to present a visual 3D model. I really should re-work that page to allow use of eg JSmol, enabled here on this blog.
♥Excepting CF3X motifs.
†FAIR data for the results reported here can be found at DOI: 10.14469/hpc/4713